Guidance for representative and targeted genomic SARS-CoV-2 monitoring
Executive Summary
Key messages
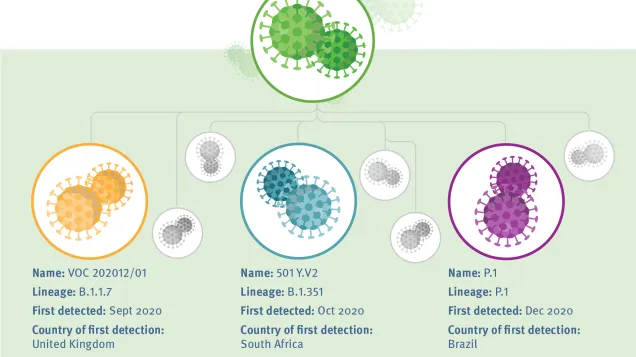
Genomic surveillance of SARS-CoV-2 is essential to detect, monitor and assess virus variants that can result in increased transmissibility, disease severity, or have other adverse effects on public health and social control measures. Obtaining timely and accurate information on the emergence and circulation of variants of concern (VOCs) and variants of interest (VOIs) requires robust surveillance systems, including integrated genome sequencing with a well-defined sampling and sequencing strategy to ensure representativeness and reliability of findings.
The current document offers practical guidance to EU/EEA Member States on implementing genomic SARS-CoV-2 surveillance, including advice on the number of samples that need to be sequenced to achieve various objectives.
For genomic surveillance of SARS-CoV-2, ECDC recommends two complementary sampling approaches:
- representative sampling of SARS-CoV-2 RT-PCR positive cases from existing, population-based surveillance systems;
- targeted sampling of SARS-CoV-2 positive cases occurring in special settings or populations.
Representative sampling and sequencing of SARS-CoV-2 cases for genomic monitoring from routine surveillance
The objective for representative sampling and sequencing of SARS-CoV-2 from cases identified during routine surveillance is to detect and monitor new variants of concern.
- For novel or emerging variant detection, ECDC recommends a detection threshold minimum which is a relative proportion of 2.5%, but ideally 1%, of a particular variant among all variants within one unit of time.
- For monitoring, ECDC recommends sequencing adequate numbers of samples to detect a difference in the relative proportion of a certain variant from one unit of time to the next (e.g. increase from 1% to 3%, or 2.5% to 5%).
Respective sample sizes and options for specimen selection are provided in the guidance.
Targeted sampling and sequencing of SARS-CoV-2 cases from special settings or populations
The following sampling frames are recommended for the respective settings or populations:
- vaccine breakthrough infections and reinfections: comprehensive sampling to detect and characterise variants causing infection in the presence of SARS-CoV-2 antibodies;
- outbreaks and clusters: a representative sample, with a minimum of five specimens per event to investigate virus transmission dynamics; detect novel genetic variants; assess the relatedness of viral strains within epidemiological clusters, and support contact tracing and other public health interventions;
- confirmed cases with travel history in areas where VOCs or VOIs are endemic: to detect potential introductions of variants and slow down their spread, ECDC recommends comprehensive sequencing sampling of all SARS-CoV-2 positive cases with travel history in areas/countries where new VOCs or VOIs are circulating;
- unusual events: a representative sample, with a minimum of five specimens from superspreading events or settings with unusually high transmission; for cases with unusual clinical presentations, ECDC recommends comprehensive sampling to support investigations of virus transmission dynamics and detection of novel genetic variants.
This guidance may need updating when the epidemiological situation changes - for example after extensive vaccination uptake in the population resulting in a significant reduction in case numbers.
Guidance for representative and targeted genomic SARS-CoV-2 monitoring
English (637.73 KB - PDF)Latest reports
Share this page